Advanced Practical Chemistry - Part I
A few months ago, I volunteered, along with Andrew McCluskey to rewrite the third-year computational chemistry lab module at the University of Bath. At the time, Andrew had just published his summer procrastination - pylj, which is an educational software that introduces students to classical atomistic simulations, and this lab seemed like the perfect opportunity to depoly it in a teaching environment.
Our course is based entirely in Jupyter Notebooks and introduces students to classical molecular dynamics using pylj, and solid state chemistry using DL_POLY and METADISE. Students will learn the theory of MD, ionic transport, defect chemsitry, and they will have the freedom to answer theiry own research questions. Over the next four weeks the students will be working through our lab course and I will write a weekly blog post, hopefully highlighting how Python is being used to enhance the learning experience of chemistry undergraduate students.
Week One
This week exclusively used pylj and involved students building, step by step their own MD code, which they then used to construct a 2D phase diagram of argon gas. Jupyter Notebooks allow us to teach in a much more interactive way. Take the Lennard-Jones model for example.
\[E_{\text{attractive}}(r_{ij}) = \frac{-B}{r_{ij}^6},\]where $B$ is some constant for the interaction, and $r_{ij}$ is the distance between the two atoms. The Pauli exclusion principle is repulsive and only present over very short distances, and therefore is modelled with the relation,
\[E_{\text{repulsive}}(r_{ij}) = \frac{A}{r_{ij}^{12}},\]again $A$ is some interaction specific constant. The total Lennard-Jones interaction is then the linear combination of these two terms,
\[E_{\text{LJ}}(r_{ij}) = E_{\text{repulsive}}(r_{ij}) + E_{\text{attractive}}(r_{ij}) = \frac{A}{r_{ij}^{12}} - \frac{B}{r_{ij}^6}.\]It is easy to present these equations and graphical representation in a lab manual, but the effectiveness of this is debatable. In our course, students were given the equation and then asked to define a function and then plot the attractive, repulsive and total energy of the Lennard-Jones interaction, as shown below.
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
def attractive(dr, b):
return -b / np.power(dr, 6)
def repulsive(dr, a):
return a / np.power(dr, 12)
def lj(dr, constants):
return repulsive(dr, constants[0]) + attractive(dr, constants[1])
r = np.linspace(3e-10, 8e-10, 100)
plt.plot(r, attractive(r, 9.273e-78), label='Attractive')
plt.plot(r, repulsive(r, 1.363e-134), label='Repulsive')
plt.plot(r, lj(r, [1.363e-134, 9.273e-78]), label='Lennard-Jones')
plt.xlabel(r'$r_{ij}$/m')
plt.ylabel(r'$E$/J')
plt.legend()
plt.show()
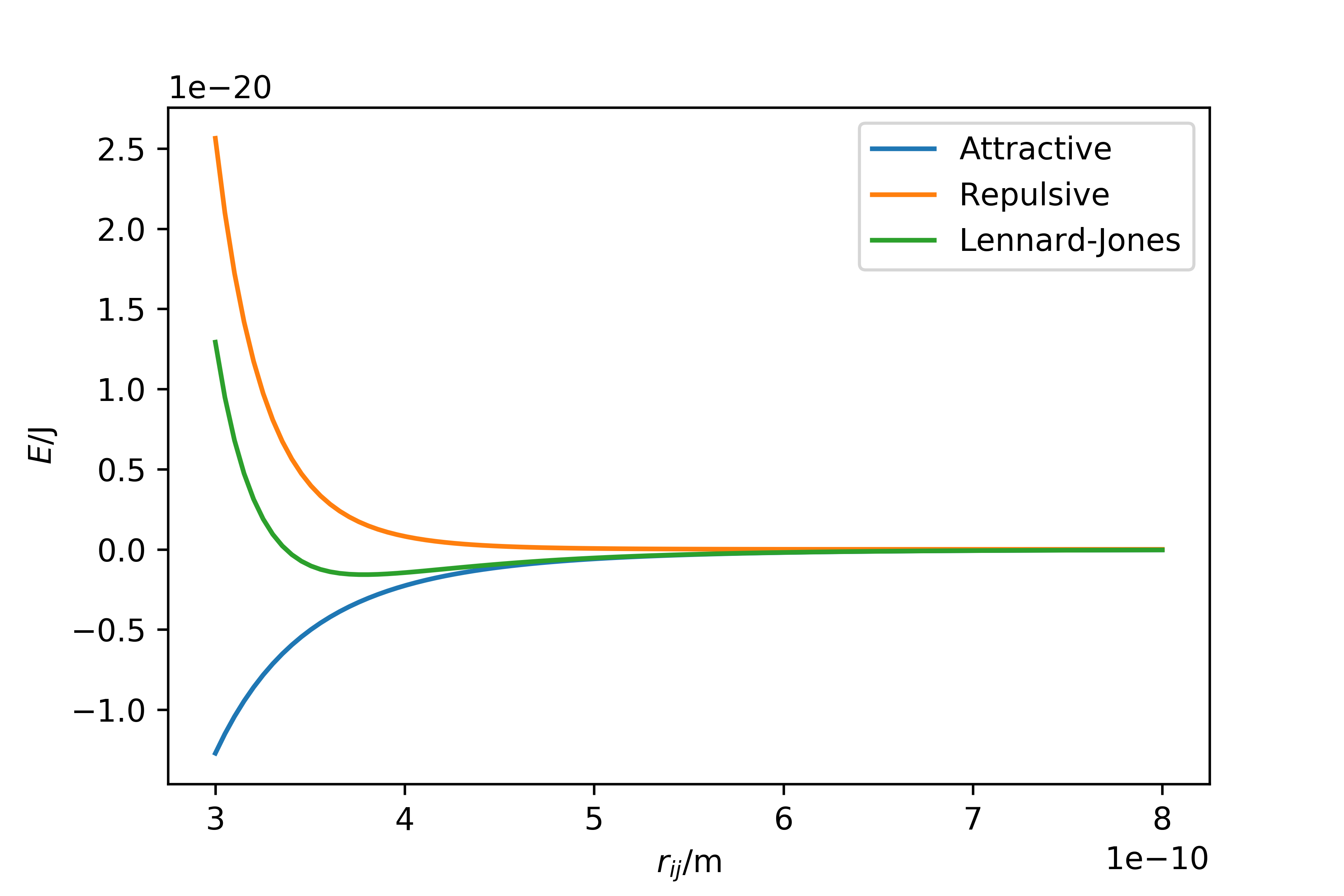
In this way, we ensured that students not only understood the Lennard-Jones potential, but also developed their programming knowledge, which at the very least is a useful transferable skill.
The first half of the course follows these lines and involves the theory of molecular dynamics written in Markdown cells within the notebook. Mixed with code cells, where students would build functions to do each part of the eventual simulation, e.g. update particle positions.
Following the introduction of MD, students had to use pylj to generate the 2D phase diagram of argon gas, by varying the temperature of the simulation and the number of particles. Below is an example of such a phase diagram.
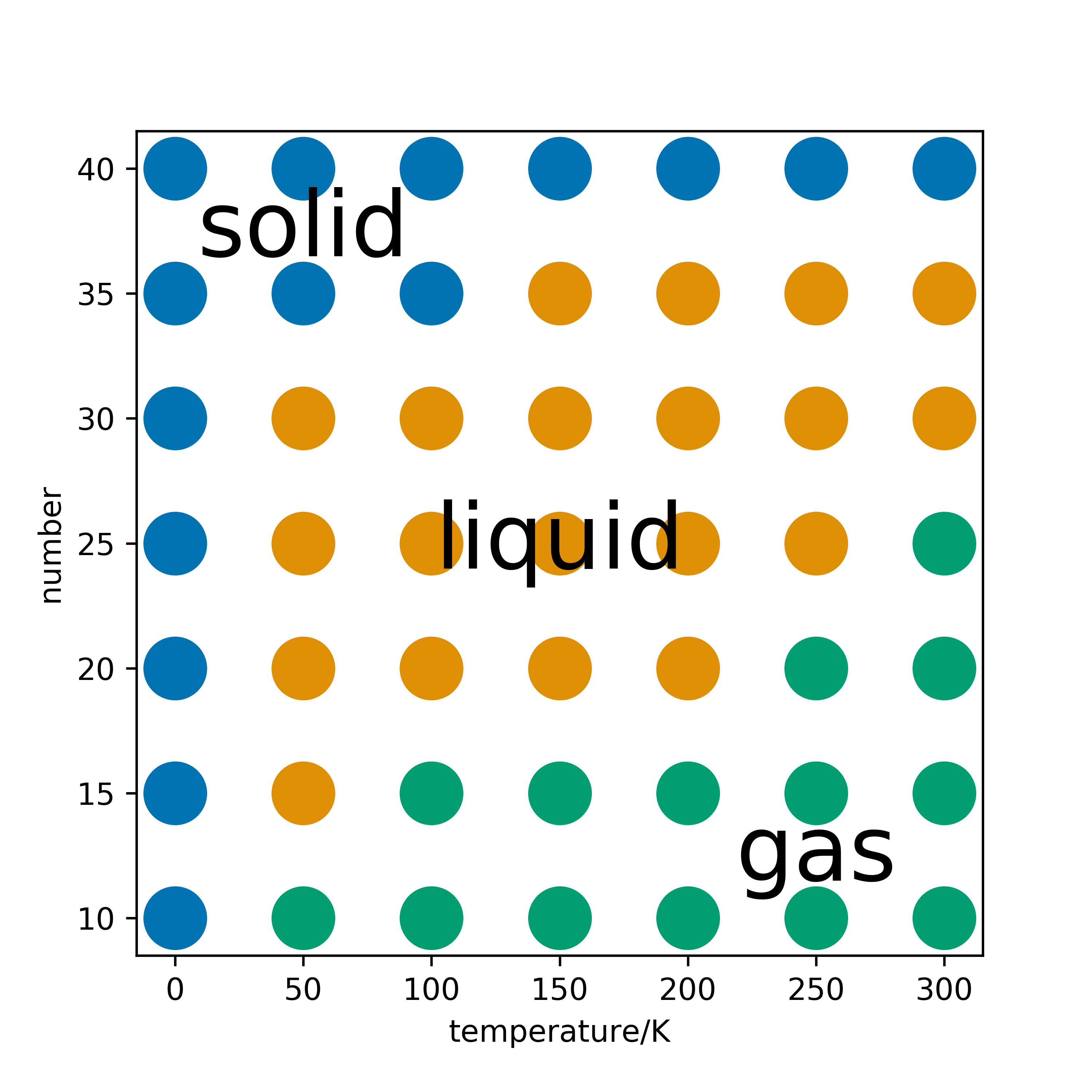
Feedback
This class contained a mix of students who had encountered Python and students who had not. There was certainly a barrier that needed to be hurdled for the non-programming students, but the lab was definitely accesible to them and they were able to achieve the learning outcomes. The non-programmers saw the trasferable skills argument and were actually very happy to try theiry hand at some light programming. There were issues with setting up the notebook before peopl ecould embark on the exercise. However, this is more down to the implementation of the Jupyter Notebooks on the University computers, than the exercise itself. In future years, more effort does need to be take to make the setup as painless as possible because it is better that students dont’ get ticked off when trying to build conda environments.
